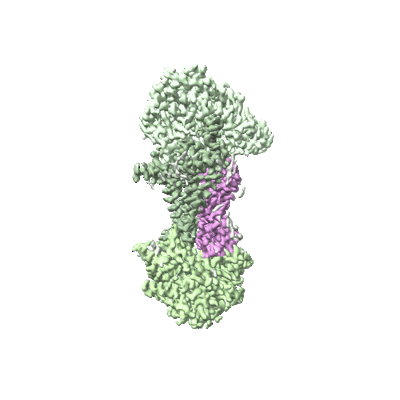
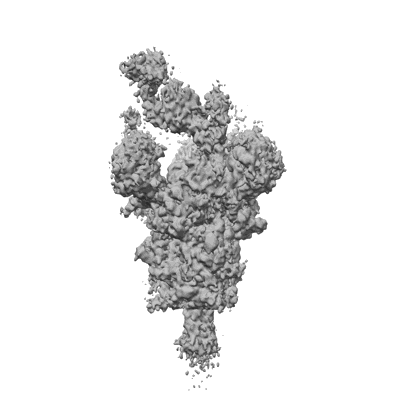
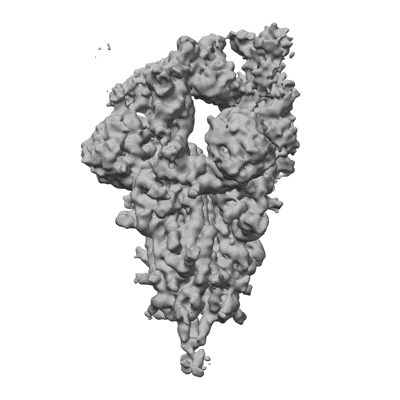Cryo-EM structure of the human parainfluenza virus hPIV3 L-P polymerase in dimeric form
Resolution: 2.7 Å
EM Method: Single-particle
Fitted PDBs: 8kdb
Q-score: 0.577
Xie J, Ouizougun-Oubari M, Wang L, Zhai G, Wu D, Lin Z, Wang M, Ludeke B, Yan X, Nilsson T, Gao L, Huang X, Fearns R, Chen S
Nat Commun (2024) 15 pp. 3163-3163 [ DOI: doi:10.1038/s41467-024-47470-7 Pubmed: 38605025 ]
- Zinc ion (65 Da, Ligand)
- Rna-directed rna polymerase l (260 kDa, Protein from Human respirovirus 3)
- Magnesium ion (24 Da, Ligand)
- Human parainfluenza virus hpiv3 l-p polymerase in dimeric form (Complex from Human respirovirus 3)
- Phosphoprotein (68 kDa, Protein from Human respirovirus 3)
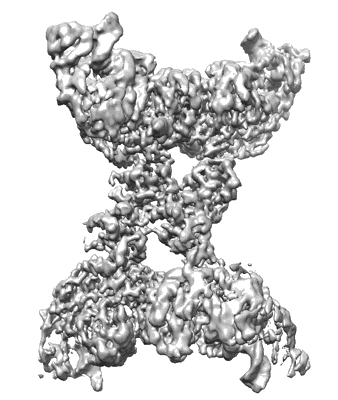
Cryo-EM structure of the human parainfluenza virus hPIV3 L-P polymerase in monomeric form
Resolution: 3.3 Å
EM Method: Single-particle
Fitted PDBs: 8kdc
Q-score: 0.506
Xie J, Ouizougun-Oubari M, Wang L, Zhai G, Wu D, Lin Z, Wang M, Ludeke B, Yan X, Nilsson T, Gao L, Huang X, Fearns R, Chen S
Nat Commun (2024) 15 pp. 3163-3163 [ DOI: doi:10.1038/s41467-024-47470-7 Pubmed: 38605025 ]
- Zinc ion (65 Da, Ligand)
- Rna-directed rna polymerase l (260 kDa, Protein from Human respirovirus 3)
- Magnesium ion (24 Da, Ligand)
- Phosphoprotein (68 kDa, Protein from Human respirovirus 3)
- Human parainfluenza virus hpiv3 l-p polymerase in monomeric form (Complex from Human respirovirus 3)
Cryo-EM structure of Mycobacterium tuberculosis OppABCD in the pre-translocation state
Resolution: 3.25 Å
EM Method: Single-particle
Fitted PDBs: 8j5q
Q-score: 0.519
Yang X, Hu T, Liang J, Xiong Z, Lin Z, Zhao Y, Zhou X, Gao Y, Sun S, Yang X, Guddat LW, Yang H, Rao Z, Zhang B
Nat Struct Mol Biol (2024) [ DOI: doi:10.1038/s41594-024-01256-z Pubmed: 38548954 ]
- Putative peptide transport permease protein rv1283c (35 kDa, Protein from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
- Iron/sulfur cluster (351 Da, Ligand)
- Uncharacterized abc transporter atp-binding protein rv1281c (65 kDa, Protein from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
- Endogenous oligopeptide (783 Da, Protein from Mycolicibacterium smegmatis MC2 155)
- Cryo-em structure of mycobacterium tuberculosis oppabcd in the pre-translocation state (Complex from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
- Putative peptide transport permease protein rv1282c (31 kDa, Protein from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
- Uncharacterized protein rv1280c (64 kDa, Protein from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
Cryo-EM structure of Mycobacterium tuberculosis OppABCD in the resting state
Resolution: 3.28 Å
EM Method: Single-particle
Fitted PDBs: 8j5r
Q-score: 0.534
Yang X, Hu T, Liang J, Xiong Z, Lin Z, Zhao Y, Zhou X, Gao Y, Sun S, Yang X, Guddat LW, Yang H, Rao Z, Zhang B
Nat Struct Mol Biol (2024) [ DOI: doi:10.1038/s41594-024-01256-z Pubmed: 38548954 ]
- Uncharacterized protein rv1280c (64 kDa, Protein from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
- Cryo-em structure of mycobacterium tuberculosis oppabcd in the resting state (Complex from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
- Putative peptide transport permease protein rv1283c (35 kDa, Protein from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
- Iron/sulfur cluster (351 Da, Ligand)
- Uncharacterized abc transporter atp-binding protein rv1281c (65 kDa, Protein from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
- Putative peptide transport permease protein rv1282c (31 kDa, Protein from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
Cryo-EM structure of Mycobacterium tuberculosis OppABCD in the pre-catalytic intermediate state
Resolution: 3.0 Å
EM Method: Single-particle
Fitted PDBs: 8j5s
Q-score: 0.566
Yang X, Hu T, Liang J, Xiong Z, Lin Z, Zhao Y, Zhou X, Gao Y, Sun S, Yang X, Guddat LW, Yang H, Rao Z, Zhang B
Nat Struct Mol Biol (2024) [ DOI: doi:10.1038/s41594-024-01256-z Pubmed: 38548954 ]
- Endogenous oligopeptide (698 Da, Protein from Mycolicibacterium smegmatis MC2 155)
- Uncharacterized abc transporter atp-binding protein rv1281c (65 kDa, Protein from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
- Putative peptide transport permease protein rv1283c (35 kDa, Protein from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
- Iron/sulfur cluster (351 Da, Ligand)
- Magnesium ion (24 Da, Ligand)
- Putative peptide transport permease protein rv1282c (31 kDa, Protein from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
- Cryo-em structure of mycobacterium tuberculosis oppabcd in the pre-catalytic intermediate state (Complex from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
- Uncharacterized protein rv1280c (64 kDa, Protein from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
- Phosphoaminophosphonic acid-adenylate ester (506 Da, Ligand)
Cryo-EM structure of Mycobacterium tuberculosis OppABCD in the catalytic intermediate state
Resolution: 2.98 Å
EM Method: Single-particle
Fitted PDBs: 8j5t
Q-score: 0.572
Yang X, Hu T, Liang J, Xiong Z, Lin Z, Zhao Y, Zhou X, Gao Y, Sun S, Yang X, Guddat LW, Yang H, Rao Z, Zhang B
Nat Struct Mol Biol (2024) [ DOI: doi:10.1038/s41594-024-01256-z Pubmed: 38548954 ]
- Uncharacterized abc transporter atp-binding protein rv1281c (65 kDa, Protein from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
- Putative peptide transport permease protein rv1283c (35 kDa, Protein from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
- Iron/sulfur cluster (351 Da, Ligand)
- Adenosine-5'-triphosphate (507 Da, Ligand)
- Magnesium ion (24 Da, Ligand)
- Cryo-em structure of mycobacterium tuberculosis oppabcd in the catalytic intermediate state (Complex from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
- Putative peptide transport permease protein rv1282c (31 kDa, Protein from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
- Uncharacterized protein rv1280c (64 kDa, Protein from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
Structure of the SPARTA complex
Resolution: 2.84 Å
EM Method: Single-particle
Fitted PDBs: 8j7s
Q-score: 0.416
Guo M, Zhu Y, Lin Z, Yang D, Zhang A, Guo C, Huang Z
Cell Res (2023) 33 pp. 731-734 [ DOI: doi:10.1038/s41422-023-00850-y Pubmed: 37491603 ]
- Dna (5'-d(p*tp*ap*ap*tp*ap*gp*ap*tp*tp*ap*gp*ap*gp*cp*cp*gp*tp*cp*ap*ap*tp*ap*gp*a)-3') (7 kDa, DNA from Maribacter polysiphoniae)
- Rna (5'-r(p*up*gp*ap*cp*gp*gp*cp*up*cp*up*ap*ap*up*cp*up*ap*up*up*a)-3') (6 kDa, RNA from Maribacter polysiphoniae)
- Tir domain-containing protein (49 kDa, Protein from Maribacter polysiphoniae)
- Piwi domain-containing protein (57 kDa, Protein from Maribacter polysiphoniae)
- Sparta complex (Complex from Maribacter polysiphoniae)
Cryo-EM structure of SARS-CoV-1 2P spike protein in complex with W328-6A1 IgG
Cryo-EM structure of SARS-CoV-1 2P spike protein in complex with W328-6E10 Fab
Cryo-EM structure of SARS-CoV-1 2P spike protein protomer in complex with W328-6E10 (local refine)